1.
Ford
D,
Easton
DF,
Stratton
M, et al.
Genetic heterogeneity and penetrance analysis of the BRCA1 and
BRCA2 genes in breast cancer families. The Breast cancer linkage
consortium . Am J Hum Genet.
1998; ; 62 :
:676.–689.
2.
Hall
JM,
Lee
MK,
Newman
B, et al.
Linkage of early-onset familial breast cancer to chromosome
17q21 . Science.
1990; ; 250 :
:1684.–1689.
3.
Miki
Y,
Swensen
J,
Shattuck-Eidens
D, et al.
A strong candidate for the breast and ovarian cancer
susceptibility gene BRCA1 . Science.
1994; ; 266 :
:66.–71.
4.
Wooster
R,
Neuhausen
SL,
Mangion
J, et al.
Localization of a breast cancer susceptibility gene, BRCA2, to
chromosome 13q12-13 . Science.
1994; ; 265 :
:2088.–2090.
5.
D’Andrea
AD,
Grompe
M. The Fanconi
anaemia/BRCA pathway . Nat Rev Cancer.
2003; ; 3 :
:23.–34.
6.
West
SC. Molecular views
of recombination proteins and their control . Nat Rev
Mol Cell Biol.
2003; ; 4 :
:435.–445.
7.
Howlett
NG,
Taniguchi
T,
Olson
S, et al.
Biallelic inactivation of BRCA2 in Fanconi
anemia . Science.
2002; ; 297 :
:606.–609.
8.
Gudmundsdottir
K,
Ashworth
A. The roles of
BRCA1 and BRCA2 and associated proteins in the maintenance of genomic
stability . Oncogene.
2006; ; 25 :
:5864.–5874.
9.
D’Andrea
A.. Susceptibility
pathways in Fanconi’s anemia and breast cancer . N
Engl J Med.
2010; ; 362 :
:1909.–1919.
10.
Roy
R,
Chun
J,
Powell
SN. BRCA1 and BRCA2:
different roles in a common pathway of genome protection .
Nat Rev Cancer.
2011; ; 12 :
:68.–78.
11.
Hakem
R, de la
Pompa
JL,
Sirard
C, et al.
The tumor suppressor gene Brca1 is required for embryonic
cellular proliferation in the mouse . Cell.
1996; ; 85 :
:1009.–1023.
12.
Patel
KJ,
Yu
VP,
Lee
H, et al.
Involvement of Brca2 in DNA repair . Mol
Cell.
1998; ; 1 :
:347.–357.
13.
Liu
J,
Doty
T,
Gibson
B,
Heyer
WD. Human BRCA2
protein promotes RAD51 filament formation on RPA-covered single-stranded
DNA . Nat Struct Mol Biol.
2010; ; 17 :
:1260.–1262.
14.
Kinzler
KW,
Vogelstein
B.
Cancer-susceptibility genes. Gatekeepers and
caretakers . Nature.
1997; ; 386 :
:761.–763.
15.
Rosen
EM,
Fan
S,
Ma
Y. BRCA1 regulation
of transcription . Cancer Lett.
2006; ; 236 :
:175.–185.
16.
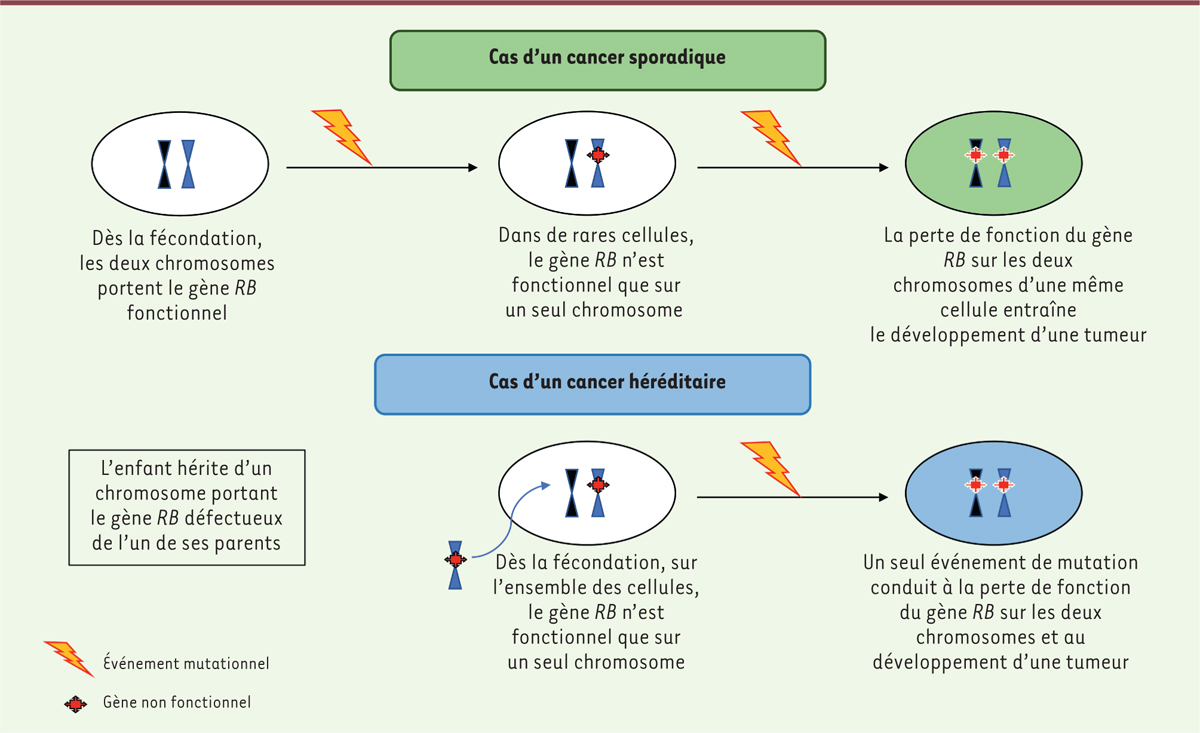
Knudson
AG. Mutation and
cancer: statistical study of retinoblastoma . Proc
Natl Acad Sci USA.
1971; ; 68 :
:820.–823.
17.
Antoniou
A,
Pharoah
PD,
Narod
S, et al.
Average risks of breast and ovarian cancer associated with BRCA1
or BRCA2 mutations detected in case series unselected for family history: a
combined analysis of 22 studies . Am J Hum
Genet.
2003; ; 72 :
:1117.–1130.
18.
King
MC,
Marks
JH,
Mandell
JB. New York Breast
Cancer Study Group. Breast and ovarian cancer risks due to inherited
mutations in BRCA1 and BRCA2 . Science.
2003; ; 302 :
:643.–646.
19.
Chen
S,
Parmigiani
G. Meta-analysis of
BRCA1 and BRCA2 penetrance . J Clin Oncol.
2007; ; 25 :
:1329.–1333.
20.
Rebbeck
TR,
Mitra
N,
Wan
F, et al.
Association of type and location of BRCA1 and BRCA2 mutations
with risk of breast and ovarian cancer .
JAMA.
2015; ; 313 :
:1347.–1361.
21.
Kuchenbaecker
KB,
McGuffog
L,
Barrowdale
D, et al.
Evaluation of polygenic risk scores for breast and ovarian cancer
risk prediction in BRCA1 and BRCA2 mutation carriers .
J Natl Cancer Inst.
2017; ; 109 :
22.
Kuchenbaecker
KB,
Hopper
JL,
Barnes
DR, et al.
Risks of breast, ovarian, and contralateral breast cancer for
BRCA1 and BRCA2 mutation carriers . JAMA.
2017; ; 317 :
:2402.–2416.
23.
Klein
AP. Genetic
susceptibility to pancreatic cancer . Mol
Carcinog.
2012; ; 51 :
:14.–24.
24.
Ginsburg
OM,
Kim-Sing
C,
Foulkes
WD, et al.
BRCA1 and BRCA2 families and the risk of skin
cancer . Fam Cancer.
2010; ; 9 :
:489.–493.
25.
Couch
FJ,
Farid
LM,
DeShano
ML, et al.
BRCA2 germline mutations in male breast cancer cases and breast
cancer families . Nat Genet.
1996; ; 13 :
:123.–125.
26.
Kote-Jarai
Z,
Leongamornlert
D,
Saunders
E, et al.
BRCA2 is a moderate penetrance gene contributing to young-onset
prostate cancer: implications for genetic testing in prostate cancer
patients . Br J Cancer.
2011; ; 105 :
:1230.–1234.
27.
Castro
E,
Goh
C,
Olmos
D, et al.
Germline BRCA mutations are associated with higher risk of nodal
involvement, distant metastasis, and poor survival outcomes in prostate
cancer . J Clin Oncol.
2013; ; 31 :
:1748.–1757.
28.
Cussenot
O.. Management of
prostate cancer: the new challenges . Presse
Med.
2017; ; 46 :
:923.–927.
29.
Moran
A,
O’Hara
C,
Khan
S, et al.
Risk of cancer other than breast or ovarian in individuals with
BRCA1 and BRCA2 mutations . Fam Cancer.
2012; ; 11 :
:235.–242.
30.
Mersch
J,
Jackson
MA,
Park
M, et al.
Cancers associated with BRCA1 and BRCA2 mutations other than
breast and ovarian . Cancer.
2015; ; 121 :
:269.–275.
31.
Eerola
H,
Heikkila
P,
Tamminen
A, et al.
Relationship of patients’age to histopathological features of
breast tumours in BRCA1 and BRCA2 and mutation-negative breast cancer
families . Breast Cancer Res.
2005; ; 7 :
:R465.–R469.
32.
Chappuis
PO,
Nethercot
V,
Foulkes
WD.
Clinico-pathological characteristics of BRCA1- and
BRCA2-related breast cancer . Semin Surg
Oncol.
2000; ; 18 :
:287.–295.
33.
Lakhani
SR,
Gusterson
BA,
Jacquemier
J, et al.
The pathology of familial breast cancer: histological features of
cancers in families not attributable to mutations in BRCA1 or
BRCA2 . Clin Cancer Res.
2000; ; 6 :
:782.–789.
34.
Foulkes
WD,
Stefansson
IM,
Chappuis
PO, et al.
Germline BRCA1 mutations and a basal epithelial phenotype in
breast cancer . J Natl Cancer Inst.
2003; ; 95 :
:1482.–1485.
35.
Foulkes
WD,
Smith
IE,
Reis-Filho
JS. Triple-negative
breast cancer . N Engl J Med.
2010; ; 363 :
:1938.–1948.
36.
Mavaddat
N,
Pharoah
PD,
Blows
F, et al.
Familial relative risks for breast cancer by pathological
subtype: a population-based cohort study . Breast
Cancer Res.
2010; ; 12 : :R10..
37.
Lakhani
SR, Van De
Vijver
MJ,
Jacquemier
J, et al.
The pathology of familial breast cancer: predictive value of
immunohistochemical markers estrogen receptor, progesterone receptor, HER-2,
and p53 in patients with mutations in BRCA1 and BRCA2 .
J Clin Oncol.
2002; ; 20 :
:2310.–2318.
38.
Bane
AL,
Beck
JC,
Bleiweiss
I, et al.
BRCA2 mutation-associated breast cancers exhibit a distinguishing
phenotype based on morphology and molecular profiles from tissue
microarrays . Am J Surg Pathol.
2007; ; 31 :
:121.–128.
39.
Agnarsson
BA,
Jonasson
JG,
Bjornsdottir
IB, et al.
Inherited BRCA2 mutation associated with high grade breast
cancer . Breast Cancer Res Treat.
1998; ; 47 :
:121.–127.
40.
Palacios
J,
Robles-Frias
MJ,
Castilla
MA, et al.
The molecular pathology of hereditary breast
cancer . Pathobiology.
2008; ; 75 :
:85.–94.
41.
Palacios
J,
Honrado
E,
Osorio
A, et al.
Immunohistochemical characteristics defined by tissue microarray
of hereditary breast cancer not attributable to BRCA1 or BRCA2 mutations:
differences from breast carcinomas arising in BRCA1 and BRCA2 mutation
carriers . Clin Cancer Res.
2003; ; 9 :
:3606.–3614.
42.
Copson
ER,
Maishman
TC,
Tapper
WJ, et al.
Germline BRCA mutation and outcome in young-onset breast cancer
(POSH): a prospective cohort study . Lancet
Oncol.
2018; ; 19 :
:169.–180.
43.
Easton
DF,
Pharoah
PD,
Antoniou
AC, et al.
Gene-panel sequencing and the prediction of breast-cancer
risk . N Engl J Med.
2015; ; 372 :
:2243.–2257.
44.
Buisson
R,
Dion-Cote
AM,
Coulombe
Y, et al.
Cooperation of breast cancer proteins PALB2 and piccolo BRCA2 in
stimulating homologous recombination . Nat Struct Mol
Biol.
2010; ; 17 :
:1247.–1254.
45.
Buisson
R,
Niraj
J,
Pauty
J, et al.
Breast cancer proteins PALB2 and BRCA2 stimulate polymerase eta
in recombination-associated DNA synthesis at blocked replication
forks . Cell Rep.
2014; ; 6 :
:553.–564.
46.
Reid
S,
Schindler
D,
Hanenberg
H, et al.
Biallelic mutations in PALB2 cause Fanconi anemia subtype FA-N
and predispose to childhood cancer . Nat
Genet.
2007; ; 39 :
:162.–164.
47.
Antoniou
AC,
Foulkes
WD,
Tischkowitz
M. Breast-cancer
risk in families with mutations in PALB2 . N Engl J
Med.
2014; ; 371 :
:1651.–1652.
48.
Casadei
S,
Norquist
BM,
Walsh
T, et al.
Contribution of inherited mutations in the BRCA2-interacting
protein PALB2 to familial breast cancer . Cancer
Res.
2011; ; 71 :
:2222.–2229.
49.
Tischkowitz
M,
Capanu
M,
Sabbaghian
N, et al.
Rare germline mutations in PALB2 and breast cancer risk: a
population-based study . Hum Mutat.
2012; ; 33 :
:674.–680.
50.
Erkko
H,
Dowty
JG,
Nikkila
J, et al.
Penetrance analysis of the PALB2 c.1592delT founder
mutation . Clin Cancer Res.
2008; ; 14 :
:4667.–4671.
51.
Heikkinen
T,
Karkkainen
H,
Aaltonen
K, et al.
The breast cancer susceptibility mutation PALB2 1592delT is
associated with an aggressive tumor phenotype . Clin
Cancer Res.
2009; ; 15 :
:3214.–3222.
52.
Thompson
ER,
Rowley
SM,
Li
N, et al.
Panel testing for familial breast cancer: calibrating the tension
between research and clinical care . J Clin
Oncol.
2016; ; 34 :
:1455.–1459.
53.
Couch
FJ,
Shimelis
H,
Hu
C, et al.
Associations between cancer predisposition testing panel genes
and breast cancer . JAMA Oncol.
2017;; 3 :
:1190.–6.
54.
Buys
SS,
Sandbach
JF,
Gammon
A, et al.
A study of over 35,000 women with breast cancer tested with a
25-gene panel of hereditary cancer genes .
Cancer.
2017; ; 123 :
:1721.–1730.
55.
Tung
N,
Battelli
C,
Allen
B, et al.
Frequency of mutations in individuals with breast cancer referred
for BRCA1 and BRCA2 testing using next-generation sequencing with a 25-gene
panel . Cancer.
2015; ; 121 :
:25.–33.
56.
Castera
L,
Krieger
S,
Rousselin
A, et al.
Next-generation sequencing for the diagnosis of hereditary breast
and ovarian cancer using genomic capture targeting multiple candidate
genes . Eur J Hum Genet.
2014; ; 22 :
:1305.–1313.
57.
Southey
MC,
Teo
ZL,
Dowty
JG, et al.
A PALB2 mutation associated with high risk of breast
cancer . Breast Cancer Res.
2010; ; 12 : :R109..
58.
Southey
MC,
Goldgar
DE,
Winqvist
R, et al.
PALB2, CHEK2 and ATM rare variants and cancer risk: data from
COGS . J Med Genet.
2016; ; 53 :
:800.–811.
59.
Ramus
SJ,
Song
H,
Dicks
E, et al.
Germline mutations in the BRIP1, BARD1, PALB2, and NBN genes in
women with ovarian cancer . J Natl Cancer
Inst.
2015; ; 107 :
60.
Rosenthal
ET,
Bernhisel
R,
Brown
K, et al.
Clinical testing with a panel of 25 genes associated with
increased cancer risk results in a significant increase in clinically
significant findings across a broad range of cancer
histories . Cancer Genet.
2017; ; 218–219 :
:58.–68.
61.
Kurian
AW,
Ward
KC,
Hamilton
AS, et al.
Uptake, results, and outcomes of germline multiple-gene
sequencing after diagnosis of breast cancer . JAMA
Oncol.
2018; ; 4 :
:1066.–1072.
62.
Cybulski
C,
Kluzniak
W,
Huzarski
T, et al.
Clinical outcomes in women with breast cancer and a PALB2
mutation: a prospective cohort analysis . Lancet
Oncol.
2015; ; 16 :
:638.–644.
63.
Jones
S,
Hruban
RH,
Kamiyama
M, et al.
Exomic sequencing identifies PALB2 as a pancreatic cancer
susceptibility gene . Science.
2009; ; 324 : :217..
64.
Moretta-Serra
J,
Berthet
P,
Bonadona
V, et al.
Recommandation française pour l’analyse en panel de gènes dans la
cadre de la prédisposition héréditaire au cancer du sein ou de l’ovaire.
Quels gènes analyser ? Pour quelle utilité clinique ? .
Bull Cancer.
2018; ; 105 :
:907.–917.
65.
Nelen
MR,
Padberg
GW,
Peeters
EA, et al.
Localization of the gene for Cowden disease to chromosome
10q22-23 . Nat Genet.
1996; ; 13 :
:114.–116.
66.
Hopkins
BD,
Parsons
RE. Molecular
pathways: intercellular PTEN and the potential of PTEN restoration
therapy . Clin Cancer Res.
2014; ; 20 :
:5379.–5383.
67.
Uppal
S,
Mistry
D,
Coatesworth
AP. Cowden disease:
a review . Int J Clin Pract.
2007; ; 61 :
:645.–652.
68.
Pilarski
R,
Stephens
JA,
Noss
R, et al.
Predicting PTEN mutations: an evaluation of Cowden syndrome and
Bannayan-Riley-Ruvalcaba syndrome clinical features .
J Med Genet.
2011; ; 48 :
:505.–512.
69.
Schrager
CA,
Schneider
D,
Gruener
AC, et al.
Clinical and pathological features of breast disease in Cowden’s
syndrome: an underrecognized syndrome with an increased risk of breast
cancer . Hum Pathol.
1998; ; 29 :
:47.–53.
70.
Bubien
V,
Bonnet
F,
Brouste
V, et al.
High cumulative risks of cancer in patients with PTEN hamartoma
tumour syndrome . J Med Genet.
2013; ; 50 :
:255.–263.
71.
Ngeow
J,
Sesock
K,
Eng
C. Breast cancer
risk and clinical implications for germline PTEN mutation
carriers . Breast Cancer Res Treat.
2017; ; 165 :
:1.–8.
72.
Li
FP,
Fraumeni
JF, Jr
,
Mulvihill
JJ, et al.
A cancer family syndrome in twenty-four kindreds .
Cancer Res.
1988; ; 48 :
:5358.–5362.
73.
Bougeard
G,
Renaux-Petel
M,
Flaman
JM, et al.
Revisiting Li-Fraumeni syndrome from TP53 mutation
carriers . J Clin Oncol.
2015; ; 33 :
:2345.–2352.
74.
Masciari
S,
Dillon
DA,
Rath
M, et al.
Breast cancer phenotype in women with TP53 germline mutations: a
Li-Fraumeni syndrome consortium effort . Breast
Cancer Res Treat.
2012; ; 133 :
:1125.–1130.
75.
Gonzalez
KD,
Noltner
KA,
Buzin
CH, et al.
Beyond Li Fraumeni syndrome: clinical characteristics of families
with p53 germline mutations . J Clin Oncol.
2009; ; 27 :
:1250.–1256.
76.
Li
J,
Meeks
H,
Feng
BJ, et al.
Targeted massively parallel sequencing of a panel of putative
breast cancer susceptibility genes in a large cohort of multiple-case breast
and ovarian cancer families . J Med Genet.
2016; ; 53 :
:34.–42.
77.
Guilford
PJ,
Hopkins
JB,
Grady
WM, et al.
E-cadherin germline mutations define an inherited cancer syndrome
dominated by diffuse gastric cancer . Hum
Mutat.
1999; ; 14 :
:249.–255.
78.
Huntsman
DG,
Carneiro
F,
Lewis
FR, et al.
Early gastric cancer in young, asymptomatic carriers of germ-line
E-cadherin mutations . N Engl J Med.
2001; ; 344 :
:1904.–1909.
79.
Fitzgerald
RC,
Hardwick
R,
Huntsman
D, et al.
Hereditary diffuse gastric cancer: updated consensus guidelines
for clinical management and directions for future research .
J Med Genet.
2010; ; 47 :
:436.–444.
80.
Pharoah
PD,
Guilford
P,
Caldas
C. International
Gastric Cancer Linkage. Incidence of gastric cancer and breast cancer in
CDH1 (E-cadherin) mutation carriers from hereditary diffuse gastric cancer
families . Gastroenterology.
2001; ; 121 :
:1348.–1353.
81.
Benusiglio
PR,
Malka
D,
Rouleau
E, et al.
CDH1 germline mutations and the hereditary diffuse gastric and
lobular breast cancer syndrome: a multicentre study .
J Med Genet.
2013; ; 50 :
:486.–489.
82.
Hansford
S,
Kaurah
P,
Li-Chang
H, et al.
Hereditary diffuse gastric cancer syndrome: CDH1 mutations and
beyond . JAMA Oncol.
2015; ; 1 :
:23.–32.
83.
Hemminki
A,
Tomlinson
I,
Markie
D, et al.
Localization of a susceptibility locus for Peutz-Jeghers syndrome
to 19p using comparative genomic hybridization and targeted linkage
analysis . Nat Genet.
1997; ; 15 :
:87.–90.
84.
Schumacher
V,
Vogel
T,
Leube
B, et al.
STK11 genotyping and cancer risk in Peutz-Jeghers
syndrome . J Med Genet.
2005; ; 42 :
:428.–435.
85.
Giardiello
FM,
Brensinger
JD,
Tersmette
AC, et al.
Very high risk of cancer in familial Peutz-Jeghers
syndrome . Gastroenterology.
2000; ; 119 :
:1447.–1453.
86.
Hearle
N,
Schumacher
V,
Menko
FH, et al.
Frequency and spectrum of cancers in the Peutz-Jeghers
syndrome . Clin Cancer Res.
2006; ; 12 :
:3209.–3215.
87.
Beggs
AD,
Latchford
AR,
Vasen
HF, et al.
Peutz-Jeghers syndrome: a systematic review and recommendations
for management . Gut.
2010; ; 59 :
:975.–986.
88.
van Lier
MG,
Wagner
A,
Mathus-Vliegen
EM, et al.
High cancer risk in Peutz-Jeghers syndrome: a systematic review
and surveillance recommendations . Am J
Gastroenterol.
2010; ; 105 :
:1258.–1264.
89.
Meindl
A,
Hellebrand
H,
Wiek
C, et al.
Germline mutations in breast and ovarian cancer pedigrees
establish RAD51C as a human cancer susceptibility gene .
Nat Genet.
2010; ; 42 :
:410.–414.
90.
Song
H,
Dicks
E,
Ramus
SJ, et al.
Contribution of germline mutations in the RAD51B, RAD51C, and
RAD51D genes to ovarian cancer in the population . J
Clin Oncol.
2015; ; 33 :
:2901.–2907.
91.
Loveday
C,
Turnbull
C,
Ramsay
E, et al.
Germline mutations in RAD51D confer susceptibility to ovarian
cancer . Nat Genet.
2011; ; 43 :
:879.–882.
92.
Sopik
V,
Akbari
MR,
Narod
SA. Genetic testing
for RAD51C mutations: in the clinic and community .
Clin Genet.
2015; ; 88 :
:303.–312.
93.
Cohen-Haguenauer
O. Quantification du
risque individuel de cancer du sein chez la femme jeune . In:
Anne Lesur
BC,
Jean-Pierre
Bellocq,
Béatrice
Gairard, eds.
La femme jeune face au cancer du sein .
Actes de la 32e Journées de la Société Française de
Sénologie et de Pathologie Mammaire; ,
Strasbourg:
2010 : :92.–107.
94.
Bell
DW,
Varley
JM,
Szydlo
TE, et al.
Heterozygous germ line hCHK2 mutations in Li-Fraumeni
syndrome . Science.
1999; ; 286 :
:2528.–2531.
95.
Shieh
SY,
Ahn
J,
Tamai
K, et al.
The human homologs of checkpoint kinases Chk1 and Cds1 (Chk2)
phosphorylate p53 at multiple DNA damage-inducible sites .
Genes Dev.
2000; ; 14 :
:289.–300.
96.
Falck
J,
Mailand
N,
Syljuasen
RG, et al.
The ATM-Chk2-Cdc25A checkpoint pathway guards against
radioresistant DNA synthesis . Nature.
2001; ; 410 :
:842.–847.
97.
Lee
JS,
Collins
KM,
Brown
AL, et al.
hCds1-mediated phosphorylation of BRCA1 regulates the DNA damage
response . Nature.
2000; ; 404 :
:201.–204.
98.
Weischer
M,
Nordestgaard
BG,
Pharoah
P, et al.
CHEK2*1100delC heterozygosity in women with breast cancer
associated with early death, breast cancer-specific death, and increased
risk of a second breast cancer . J Clin
Oncol.
2012; ; 30 :
:4308.–4316.
99.
Wang
N,
Ding
H,
Liu
C, et al.
A novel recurrent CHEK2 Y390C mutation identified in high-risk
Chinese breast cancer patients impairs its activity and is associated with
increased breast cancer risk . Oncogene.
2015; ; 34 :
:5198.–5205.
100.
Savitsky
K,
Bar-Shira
A,
Gilad
S, et al.
A single ataxia telangiectasia gene with a product similar to
PI-3 kinase . Science.
1995; ; 268 :
:1749.–1753.
101.
Cavaciuti
E,
Lauge
A,
Janin
N, et al.
Cancer risk according to type and location of ATM mutation in
ataxia-telangiectasia families . Genes Chromosomes
Cancer.
2005; ; 42 :
:1.–9.
102.
Thompson
D,
Duedal
S,
Kirner
J, et al.
Cancer risks and mortality in heterozygous ATM mutation
carriers . J Natl Cancer Inst.
2005; ; 97 :
:813.–822.
103.
Olsen
JH,
Hahnemann
JM,
Borresen-Dale
AL, et al.
Breast and other cancers in 1445 blood relatives of 75 Nordic
patients with ataxia telangiectasia . Br J
Cancer.
2005; ; 93 :
:260.–265.
104.
d’Almeida
AK,
Cavaciuti
E,
Dondon
MG, et al.
Increased risk of breast cancer among female relatives of
patients with ataxia-telangiectasia: a causal relationship?
Br J Cancer.
2005;; 93 :
:730.–2.
105.
Renwick
A,
Thompson
D,
Seal
S, et al.
ATM mutations that cause ataxia-telangiectasia are breast cancer
susceptibility alleles . Nat Genet.
2006; ; 38 :
:873.–875.
106.
Balleine
RL,
Murali
R,
Bilous
AM, et al.
Histopathological features of breast cancer in carriers of ATM
gene variants . Histopathology.
2006; ; 49 :
:523.–532.
107.
Tavtigian
SV,
Oefner
PJ,
Babikyan
D, et al.
Rare, evolutionarily unlikely missense substitutions in ATM
confer increased risk of breast cancer . Am J Hum
Genet.
2009; ; 85 :
:427.–446.
108.
Goldgar
DE,
Healey
S,
Dowty
JG, et al.
Rare variants in the ATM gene and risk of breast
cancer . Breast Cancer Res.
2011; ; 13 : :R73..
109.
Chenevix-Trench
G,
Spurdle
AB,
Gatei
M, et al.
Dominant negative ATM mutations in breast cancer
families . J Natl Cancer Inst.
2002; ; 94 :
:205.–215.
110.
Stankovic
T,
Kidd
AM,
Sutcliffe
A, et al.
ATM mutations and phenotypes in ataxia-telangiectasia families in
the British Isles: expression of mutant ATM and the risk of leukemia,
lymphoma, and breast cancer . Am J Hum Genet.
1998; ; 62 :
:334.–345.
111.
Roberts
NJ,
Jiao
Y,
Yu
J, et al.
ATM mutations in patients with hereditary pancreatic
cancer . Cancer Discov.
2012; ; 2 :
:41.–46.
112.
Helgason
H,
Rafnar
T,
Olafsdottir
HS, et al.
Loss-of-function variants in ATM confer risk of gastric
cancer . Nat Genet.
2015; ; 47 :
:906.–910.
113.
Kurian
AW,
Hare
EE,
Mills
MA, et al.
Clinical evaluation of a multiple-gene sequencing panel for
hereditary cancer risk assessment . J Clin
Oncol.
2014; ; 32 :
:2001.–2009.
114.
De Nicolo
A,
Tancredi
M,
Lombardi
G, et al.
A novel breast cancer-associated BRIP1 (FANCJ/BACH1) germ-line
mutation impairs protein stability and function .
Clin Cancer Res.
2008; ; 14 :
:4672.–4680.
115.
Eisinger
F,
Bressac
B,
Castaigne
D, et al.
Identification and management of hereditary predisposition to
cancer of the breast and the ovary (update 2004) .
Bull Cancer.
2004; ; 91 :
:219.–237.
116.
Szabo
C,
Masiello
A,
Ryan
JF,
Brody
LC. The breast
cancer information core: database design, structure, and
scope . Hum Mutat.
2000; ; 16 :
:123.–131.
117.
Beroud
C,
Letovsky
SI,
Braastad
CD, et al.
BRCA Share: a collection of clinical BRCA gene
variants . Hum Mutat.
2016; ; 37 :
:1318.–1328.
118.
King
MC,
Marks
JH,
Mandell
JB. New York Breast
Cancer Study G. Breast and ovarian cancer risks due to inherited mutations
in BRCA1 and BRCA2 . Science.
2003; ; 302 :
:643.–646.
119.
Muller
D,
Bonaiti-Pellie
C,
Abecassis
J, et al.
BRCA1 testing in breast and/or ovarian cancer families from
northeastern France identifies two common mutations with a founder
effect . Fam Cancer.
2004; ; 3 :
:15.–20.
120.
Plon
SE,
Eccles
DM,
Easton
D, et al.
Sequence variant classification and reporting: recommendations
for improving the interpretation of cancer susceptibility genetic test
results . Hum Mutat.
2008; ; 29 :
:1282.–1291.
121.
Spurdle
AB,
Healey
S,
Devereau
A, et al.
ENIGMA: evidence-based network for the interpretation of germline
mutant alleles: an international initiative to evaluate risk and clinical
significance associated with sequence variation in BRCA1 and BRCA2
genes . Hum Mutat.
2012; ; 33 :
:2.–7.
122.
Rebbeck
TR,
Friebel
TM,
Friedman
E, et al.
Mutational spectrum in a worldwide study of 29,700 families with
BRCA1 or BRCA2 mutations . Hum Mutat.
2018; ; 39 :
:593.–620.